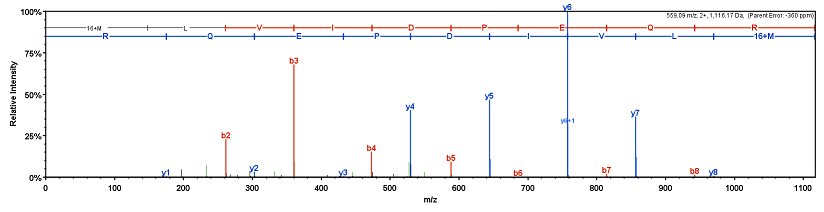
Performance Investigation of Proteomic Identification by HCD/CID Fragmentations in Combination with High/Low-Resolution Detectors on a Tribrid, High-Field Orbitrap Instrument | PLOS ONE
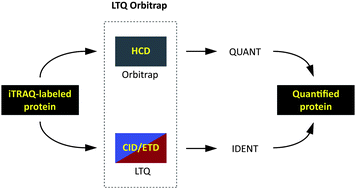
Gaining efficiency by parallel quantification and identification of iTRAQ-labeled peptides using HCD and decision tree guided CID/ETD on an LTQ Orbitrap - Analyst (RSC Publishing)

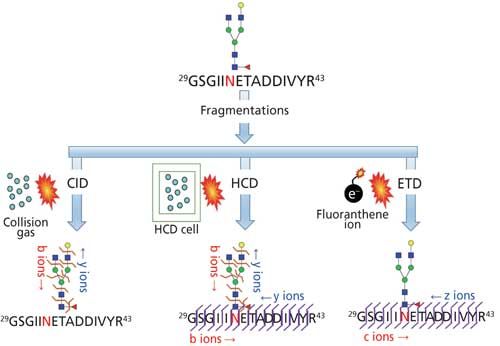
Higher Energy Collision Dissociation (HCD) Product Ion-Triggered Electron Transfer Dissociation (ETD) Mass Spectrometry for the Analysis of N-Linked Glycoproteins | Journal of Proteome Research

Low‐energy collision‐induced dissociation (low‐energy CID), collision‐induced dissociation (CID), and higher energy collision dissociation (HCD) mass spectrometry for structural elucidation of saccharides and clarification of their dissolution ...

CID/HCD MS 3 spectrum of m/z 741.36, i.e. y9 ion resulting from CID... | Download Scientific Diagram
![PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d76885d896eb57c5dcb781600ed39724f4c46016/7-Figure4-1.png)
PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar

Expediting SRM Assay Development for Large-Scale Targeted Proteomics Experiments | Journal of Proteome Research
![PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d76885d896eb57c5dcb781600ed39724f4c46016/6-Figure3-1.png)
PDF] Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics* | Semantic Scholar
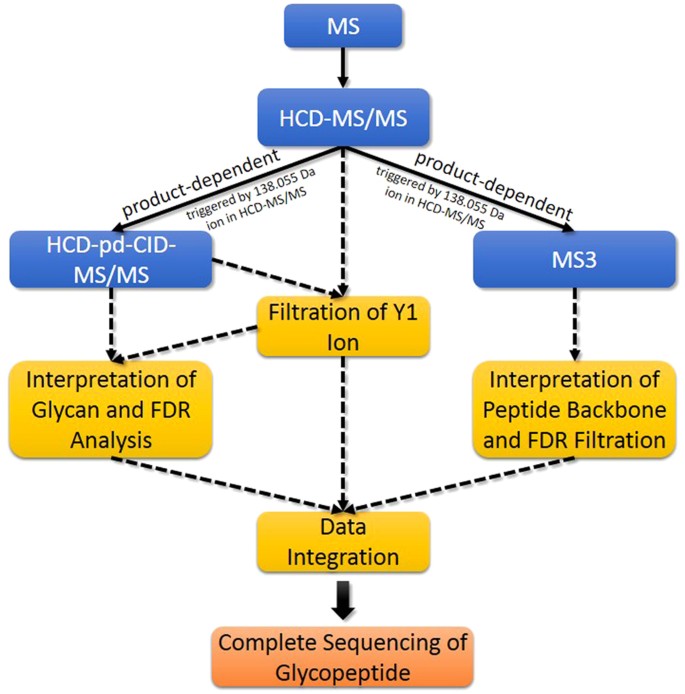
pGlyco: a pipeline for the identification of intact N-glycopeptides by using HCD- and CID-MS/MS and MS3 | Scientific Reports

Current algorithmic solutions for peptide-based proteomics data generation and identification. - Abstract - Europe PMC

Characterization of collision-induced dissociation of deprotonated peptides of 4–16 amino acids using high-resolution mass spectrometry - ScienceDirect
Performance Investigation of Proteomic Identification by HCD/CID Fragmentations in Combination with High/Low-Resolution Detectors on a Tribrid, High-Field Orbitrap Instrument | PLOS ONE

Table 1 from Improving collision induced dissociation (CID), high energy collision dissociation (HCD), and electron transfer dissociation (ETD) fourier transform MS/MS degradome-peptidome identifications using high accuracy mass information. | Semantic ...

Developments, advancements, and contributions of mass spectrometry in omics technologies - ScienceDirect

Finding and using diagnostic ions in collision induced crosslinked peptide fragmentation spectra - ScienceDirect

Evaluation of HCD- and CID-type Fragmentation Within Their Respective Detection Platforms For Murine Phosphoproteomics - ScienceDirect